|
Plot(booteval.relimp(boot,sort=TRUE)) # plot result 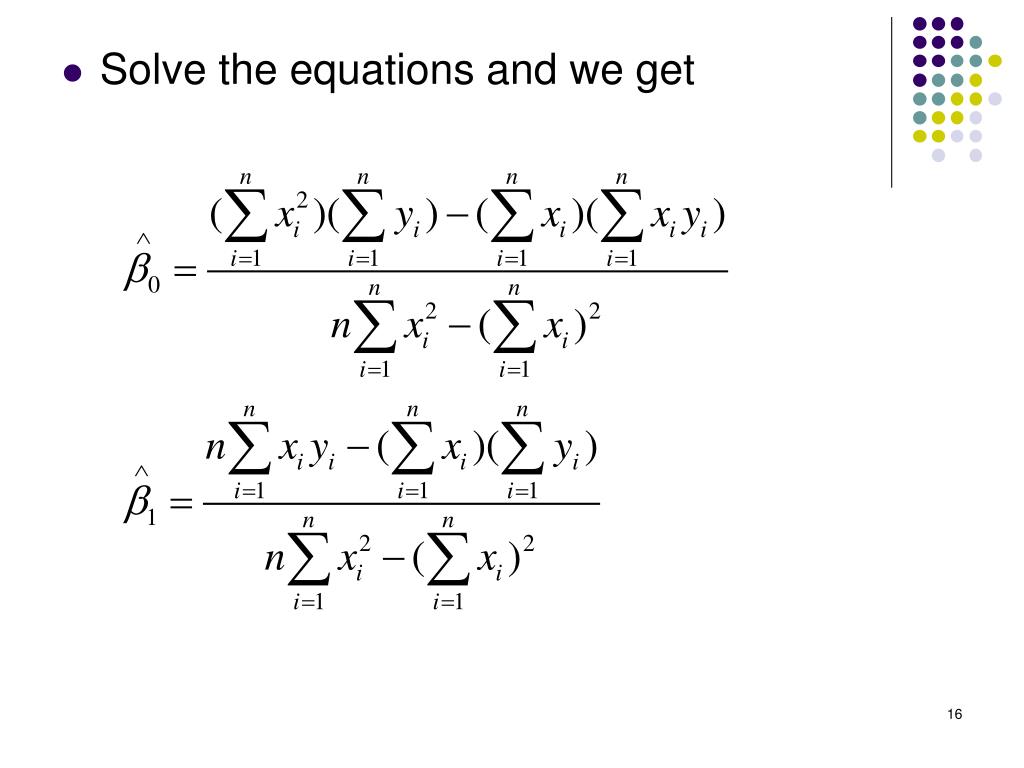
# Bootstrap Measures of Relative Importance (1000 samples)īoot <- boot.relimp(fit, b = 1000, type = c("lmg", # Calculate Relative Importance for Each PredictorĬalc.relimp(fit,type=c("lmg","last","first","pratt"), See help(calc.relimp) for details on the four measures of relative importance provided. The relaimpo package provides measures of relative importance for each of the predictors in the model. Other options for plot( ) are bic, Cp, and adjr2. Models are ordered by the selection statistic. # plot a table of models showing variables in each model. Here, the ten best models will be reported for each subset size (1 predictor, 2 predictors, etc.). In the following code nbest indicates the number of subsets of each size to report. stepAIC( ) performs stepwise model selection by exact AIC.Īlternatively, you can perform all-subsets regression using the leaps( ) function from the leaps package. You can perform stepwise selection (forward, backward, both) using the stepAIC( ) function from the MASS package. Selecting a subset of predictor variables from a larger set (e.g., stepwise selection) is a controversial topic. Results <- crossval(X,y,theta.fit,theta.predict,ngroup=10)Ĭor(y,results$cv.fit)**2 # cross-validated R2 Variable Selection # Assessing R2 shrinkage using 10-Fold Cross-Validation Using the crossval() function from the bootstrappackage, do the following: You can assess R2 shrinkage via K-fold cross-validation. Sum the MSE for each fold, divide by the number of observations, and take the square root to get the cross-validated standard error of estimate. You can do K-Fold cross-validation using the cv.lm( ) function in the DAAG package.Ĭv.lm(df=mydata, fit, m=3) # 3 fold cross-validation The following code provides a simultaneous test that x3 and x4 add to linear prediction above and beyond x1 and x2.įit1 <- lm(y ~ x1 + x2 + x3 + x4, data=mydata) You can compare nested models with the anova( ) function. Layout(matrix(c(1,2,3,4),2,2)) # optional 4 graphs/pageįor a more comprehensive evaluation of model fit see regression diagnostics or the exercises in this interactive course on regression. Vcov(fit) # covariance matrix for model parametersĭiagnostic plots provide checks for heteroscedasticity, normality, and influential observerations. Fitting the ModelĬonfint(fit, level=0.95) # CIs for model parameters The topics below are provided in order of increasing complexity. The two functions that can be used to visualize a linear fit are regplot() and lmplot().R provides comprehensive support for multiple linear regression. Functions for drawing linear regression models # The goal of seaborn, however, is to make exploring a dataset through visualization quick and easy, as doing so is just as (if not more) important than exploring a dataset through tables of statistics. To obtain quantitative measures related to the fit of regression models, you should use statsmodels. That is to say that seaborn is not itself a package for statistical analysis. In the spirit of Tukey, the regression plots in seaborn are primarily intended to add a visual guide that helps to emphasize patterns in a dataset during exploratory data analyses. The functions discussed in this chapter will do so through the common framework of linear regression. 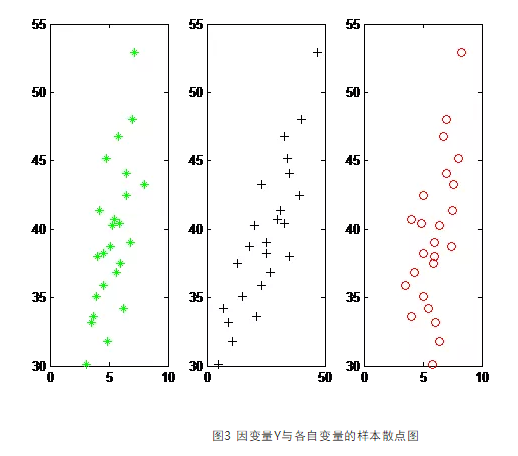
It can be very helpful, though, to use statistical models to estimate a simple relationship between two noisy sets of observations. We previously discussed functions that can accomplish this by showing the joint distribution of two variables. Many datasets contain multiple quantitative variables, and the goal of an analysis is often to relate those variables to each other.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
